
Andreas Hilfinger
-
E-mail:
-
Website:
-
Mailing Address:
3359 Mississauga Road
Mississauga ON L5L 1C6
Canada
Research Areas:
Theoretical Biophysics; Stochastic models of complex cellular processes
Research Profile:
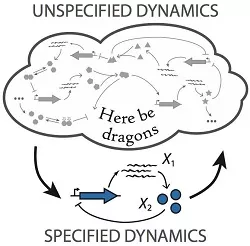
One of the key challenges of understanding biological cells is trying to make rigorous and quantitative statements in the face of enormous model uncertainty: While we typically know some things about a cellular process we never know everything, which leaves mathematical models hugely under-determined and makes common modeling approaches unreliable.
The focus of our research is thus to study complex systems such as cellular processes, not by ignoring or guessing unknown details, but by deriving testable predictions that are provably independent of them. For example, specifying some features of a system while leaving everything else unspecified allows us to establish physical performance bounds for classes of intracellular processes such as feedback control.
Additionally, we develop theoretical tools to exploit naturally occurring cell-to-cell variability to test specific hypotheses within large reaction networks. For example, the network invariants we derived showed that mRNA-protein fluctuation data in E. coli contradict the majority of published models of stochastic gene expression.
Courses Taught:
PHY147H5, PHY325H5, and PHY451H5 (undergraduate); PHY2707H1 (graduate)
Publications
Jager K, Orozco-Hildago MT, Springstein BL, Joly-Smith E, Papazotos F, McDonough E, Fleming E,
McCallum G, Hilfinger A, Hochschild A, and Potvin-Trottier L. Measuring prion propagation in single bacteria elucidates mechanism of loss. Proceedings of the National Academy of Sciences 120 (39) e2221539120 (2023)
Kell B, Ripsman R, and Hilfinger A. Noise properties of adaptation-conferring biochemical control modules. Proceedings of the National Academy of Sciences 120 (38) e2302016120 (2023)
Diep P, Kell B, Yakunin A, Hilfinger A, and Mahadevan R. Quantifying metal-binding specificity of CcNikZ-II from Clostridium carboxidivorans in the presence of
competing metal ions. Analytical Biochemistry 676, 115182, (2023)
Fan, R & Hilfinger A. The effect of microRNA on protein variability and gene expression fidelity. Biophysical Journal 122, 1–19 (2023)
T Wittenstein, N Leibovich, A Hilfinger. Quantifying biochemical reaction rates from static population variability within incompletely observed complex networks. PLoS Computational Biology 18.6, (2022)
T Kaufman, E Nitzan, N Firestein, MB Ginzberg, S Iyengar, N Patel, R Ben-Hamo, Z Porat, J Hunter, A Hilfinger, V Rotter, R Kafri, R Straussman. Visual barcodes for multiplexing live microscopy-based assays. Nature Communications, 13, 2725, (2022)
Joly-Smith, E., Wang, Z. J., Hilfinger, A. Inferring gene regulation dynamics from static snapshots of gene expression variability. Physical Review E (2021), 104(4), 044406
Ta, C., Ginzberg, M.B., Webster, R., Iyengar, S., Liu, S., Concannon, J., Wang, Y., Auld, D.S., Jenkins, J.L., Rost, H., Hilfinger, A., Derry, W.B., Patel, N., Kafri, R. Cell size homeostasis is maintained bt a circuitry involving a CDK4-determined target size that programs the cell size-dependent action of p38. Developmental Cell (2021), June, 56, 12, P1756-1769.E7
Yan, J., Hilfinger, A., Vinnicombe, G. and Paulsson, J. (2019). Kinetic Uncertainty Relations for the Control of Stochastic Reaction Networks, Physical Review Letters, 123 (10), 108101.
Hilfinger, A., Norman, T.M., and Paulsson, J. (2016). Exploiting Natural Fluctuations to Identify Kinetic Mechanisms in Sparsely Characterized Systems, Cell Systems, 2, 251–259.
Hilfinger, A., Norman, T.M., Vinnicombe, G., and Paulsson, J. (2016). Constraints on Stochastic Fluctuations in Sparsely Characterized Biological Systems, Physical Review Letters, 116, 058101.
Hilfinger, A., Chen, M., and Paulsson, J. (2012). Using Temporal Correlations and Full Distributions to Separate Intrinsic and Extrinsic Fluctuations In Dynamic Biological Systems, Physical Review Letters, 109, 248104.
Hilfinger, A. and Paulsson, J. (2011). Separating Intrinsic from Extrinsic Fluctuations in Dynamic Biological Systems, Proc. of the National Academy of Sciences, 108(29), 12167-12172.
Hilfinger, A., Chattopadhyay, A.K., and Jülicher, F. (2009). Nonlinear Dynamics of Cilia and Flagella, Physical Review E, 79 (5 Pt 1), 051918.
Hilfinger, A. and Jülicher, F. (2008). The Chirality of Ciliary Beats, Physical Biology, 5(1), 16003.
Riedel-Kruse, I.H., Hilfinger, A., Howard, J., and Jülicher, F. (2007). How Molecular Motors Shape the Flagellar Beat, Human Frontiers Science Program (HFSP) Journal, 1(3), 192-208.
