
Patrick Gunning
-
E-mail:
-
Phone:
-
Fax:
-
Website:
-
Mailing Address:
3359 Mississauga Road
Mississauga ON L5L 1C6
Canada
Research Areas:
Medicinal Chemistry, Biotechnology, Chemosessors, Cancer Biology, Rare Disease Therapeutics
Research Profile:
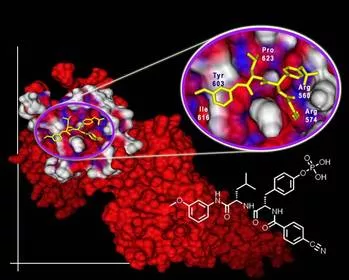
Most biological processes involve permanent and nonpermanent interactions between different proteins. Many protein complexation events play key roles in various human diseases. The thrust of our research focuses on the development of novel, small molecule architectures to reverse a protein's aberrant role, via manipulation of protein complexation events. Numerous protein-protein interfaces contain compact, centralized regions of residues, known as 'hot spots', crucial for interaction. Many proteins function by binding to multiple partners and these proteins tend to use the same hot spots, which can adapt to present the same residues in different structural contexts.
Our research seeks to design and develop scaffolds to artificially suppress or up-regulate specific gene expression profiles via manipulation of protein-protein interactions, thereby inducing therapeutically-beneficial cellular responses in malfunctioning human cells. The proposed research seeks to validate whether protein function can be 'switched' on or off through artificial protein complexation by divalent conjugated small molecule 'hot spot' recognition agents. Molecular modulation of specific protein-protein interactions offers a dynamic approach to artificially regulating aberrant protein activity in human disease. A keen objective of the proposed work is to promote and illuminate the efficacy of developing novel drug-like scaffolds incorporating inorganic, as well as organic, functionality to achieve in vivo manipulation of cellular signaling.
Courses Taught:
CHM243H5, CHM345H5, CHM444H5, and CPS400Y5 (undergraduate); CHM1009H1 and CHM1051H1 (graduate)
Publications
Recent:
Abdallah, Diaaeldin I., Elvin D. de Araujo, Naman H. Patel, Lina S. Hasan, Richard Moriggl, Oliver H. Krämer, and Patrick T. Gunning. (2023). Medicinal chemistry advances in targeting class I histone deacetylases. Exploration of Targeted Anti-tumor Therapy 4, no. 4757.
de Araujo, E. D., Orlova, A., Ashraf, Q. F., Moriggl, R., & Gunning, P. T. (2023). Inhibitor Library Screening of SH2 Domains Through Denaturation-Based Assays. In SH2 Domains: Functional Modules and Evolving Tools in Biology. 213-223. New York, NY: Springer US. D
Geordon A. Frere, Advait Hasabnis, Camila B. Francisco, Motasem Suleiman, Olga Alimowska, Rima Rahmatullah, Jerome Gould, Celia Yi-Chia Su, Oleksandr Voznyy, Patrick T. Gunning, Ernani A. Basso, and Robert S. Prosser (2024 Next-Generation Tags for Fluorine Nuclear Magnetic Resonance: Designing Amplification of Chemical Shift Sensitivity. Journal of the American Chemical Society.146 (5), 3052-3064
L. Metzelder, Michael Bergmann, Maik Dahlhoff, Florian Grebien, Roman Fleck, Christine Pirker, Walter Berger, Emir Hadzijusufovic, Wolfgang R. Sperr, Lukas Kenner, Peter Valent, Tero Aittokallio, Marco Herling, Satu Mustjoki, Patrick T. Gunning, Richard Moriggl (2023). Small molecule STAT3/5 inhibitors exhibit therapeutic potential in acute myeloid leukemia and extra-nodal natural killer/T cell lymphoma. 134(8)
Yasir S. Raouf, Abootaleb Sedighi, Mulu Geletu, Geordon A. Frere, Rebecca G. Allan, Nabanita Nawar, Elvin D. de Araujo, and Patrick T. Gunning (2023). Discovery of YSR734: A Covalent HDAC Inhibitor with Cellular Activity in Acute Myeloid Leukemia and Duchenne Muscular Dystrophy. Journal of Medicinal Chemistry. 66 (24), 16658-16679.
Yasir S. Raouf, Abootaleb Sedighi, Mulu Geletu, Geordon A. Frere, Rebecca G. Allan, Nabanita Nawar, Elvin D. de Araujo, and Patrick T. Gunning (2023). Discovery of YSR734: A Covalent HDAC Inhibitor with Cellular Activity in Acute Myeloid Leukemia and Duchenne Muscular Dystrophy. Journal of Medicinal Chemistry. 66 (24), 16658-16679.
Hadzijusufovic E., Keller A., Berger D., Greiner G., Wingelhofer B., Witzeneder N., Ivanov D., Pecnard E., Nivarthi H., Schur F.K.M., Filik Y., Kornauth C., Neubauer H.A., Mullauer L., Tin G., Park, J., de Araujo E.D., Gunning P.T., Hoermann G., Gouilleux F., Kralovics R., Moriggl R., Valent P. (2020). STAT5B is expressed in CD34+/CD38- stem cells and serves as a potential molecular target in Phnegative myeloproliferative neoplasms. Cancers. 12:1021-1029
Abdeldayem, A., Raouf, Y. S., Constantinescu, S., Moriggl, R., Gunning, P. T. (2020). “Advances in Covalent Kinase Inhibitors.” ChemSocRev. 49:2617-2687.
Manaswiyoungkul P., Erdogan F., Olaoye O. O., de Araujo E. D., Gunning P. T. (2020). “The Development and Optimization of STAT5B High-Throughput Fluorescence Polarization Assay for DNA-Binding Domain-Targeting Inhibitors.” J. Pharm Biomed Anal, 184: 113182-113189.
Shouksmith A. E., Gawel J. M., Nawar N., Sina D., Raouf Y. S., Bukhari S., He L., Johns A. E., Manaswiyoungkul P., Olaoye O. O., Cabral A. D., Sedighi A., de Araujo E. D., Gunning P.T. (2020). Class I/IIb-Selective HDAC Inhibitor Exhibits Oral Bioavailability and Therapeutic Efficacy in Acute Myeloid Leukemia. ACS Med Chem Lett., 11:56-64.
Maurer B., Nivarthi H., Wingelhofer B., Pham H. T. T., Grausenburger R., Schlederer M., Prchal-Murphy M., Chen D., Winkler S., Merkel O., Schiefer A-I., Kornauth C., Suske T., Hofbauer M., Hochgatterer B., Hoermann G., Hoelbl-Kovacic A., Cumaraswamy A. A., Eder G., Kitzwögerer M., Chott A., Pospíšilova S., Kubicek S., Valent P., Kolbe T., Grebien T., Kenner L., Gunning P. T., Kralovics R., Herling M., Müller M., Rülicke T., Sexl V., Moriggl R. (2020). “High activation of STAT5A drives Peripheral T-Cell Lymphoma and Leukemia” Haematologica, 105, 435-447.
Geletu M., Hoskin V., Niit M., Adan H., Gunning P. T., Raptis L. (2019). “Differentiation of mouse breast epithelial HC11 and EPH4 cells", JoVE, JoVE60147
Manaswiyoungkul P., de Araujo E. D., and Gunning P.T. (2019). “Targeting Prenylation Inhibition through the Mevalonate Pathway.” RSC Med Chem., 11, 51-71.
de Araujo E.D., Orlova A., Neubauer H.A., Bajusz D., Seo H-S., Dhe-Paganon S., Keserű G.M., Moriggl R., Gunning P.T. (2019). Structural Implications of STAT3 and STAT5 SH2 Domain Mutations. Cancers. 11, 1757-1778.
Orlova A., Wagner C., de Araujo E. D., Bajusz D., Neubauer H. A., Herling M., Gunning P. T., Keserű G. M., Morrigl, R. (2019). Direct targeting options of STAT3 and STAT5 in cancer. Cancers, 11, 1930-1946.
Nitt, M., Geletu M., Taha Z., Arulanandam R., Cass J., Hoskin V., Elliot B., Gunning P. T., Raptis L. (2019). “Regulation of Differentiation of HC11 Mouse Breast Epithelial Cells by the Signal Transducer and Activator of Transcription-3” Anticancer Research, 39, 2749-2756.
Geletu, M., Taha, Z., Arulanandam, R., Mohan, R., Assi, H. H., Castro, M. G., Nabi, I. R., Gunning, P. T., Raptis, L. (2019). “Effect of Caveolin-1 upon Stat3-pTyr705 levels in breast and lung carcinoma cells.” Biochem Cell Biol., 97, 638-646.
de Araujo E. D., Erdogan F., Neubauer H. A., Meneksedag-Erol D., Manaswiyoungkul P., Eram M. S., Seo H-S., Qadree A. K., Israelian J., Orlova A., Suske T., Pham H. T. T., Boersma A., Tangermann S., Kenner L., Rülicke T., Dong A., Ravichandran M., Brown P. J., Audette G. F., Rauscher S., Dhe-Paganon S., Moriggl R., and Gunning P.T. (2019).* “Structural and Functional Consequences of the STAT5BN642H Driver Mutation.” Nature Commun., 10, 1-15.
Linher-Melville K., Sharma M., Nakhla P., Kum E., Ungard R., Park J., Rosa D.A., Gunning P.T., Singh G. (2019). “Inhibiting STAT3 in a murine model of human breast cancer-induced bone pain delays the onset of nociception.” Molecular Pain, 15, 1-16.
de Araujo E. D., Manaswiyoungkul P., Erdogan F., Qadree A. K., Sina D., Tin G., Toutah K., Yuen K., Gunning P. T. (2019). “A functional in vitro assay for screening inhibitors of STAT5B phosphorylation” J. Pharm Biomed Anal, 162, 60-65.
Geletu M., Taha Z., Gunning P. T., Raptis L. (2019). “PI3k and Stat3: Oncogenes that are Required for Gap Junctional, Intercellular Communication.” Cancers, 11, 167.
Kosack L., Wingelhofer B., Popa A., Orlova a., Agerer B., Vilagos B., Majek P., Parapatics K., Lercher A., Ringler A., Klughammer J., Smyth M., Khamina K., Baazim H., de Araujo E. D., Rosa D. A., Park J., Tin G., Ahmar S., Gunning P. T., Bock C, Siddle H.V., Woods G. M., Kubicek S., Murchison E. P., Bennett K. L., Moriggl R., Bergthaler A. (2019).* “The ERBB-STAT3 axis drives Tasmanian devil facial tumor disease.” Cancer Cell, 35, 125-139.
Shouksmith, A. E., Grimard, M. L., Geletu, M., de Araujo, E. D., Berger-Becvar, A., Heaton, W. L., Gaweł, J. L., Bakhshinyan, D., Adile, A. A., Venugopal, C., Johns, A. E., Al-Qaysi, O., Lewis, A. M., O’Hare, T., Deininger, M., Singh, S. K., Luchman, A., Weiss, S., Fishel, M. L.,* and Gunning, P. T. (2019).* “Identification and characterization of AES-135, a potent HDAC inhibitor that prolongs survival in an orthotopic mouse model of pancreatic cancer. J. Med. Chem., 62, 2651-2665.
Demarez, C., Gérard, C., Cordi, S., Poncy, A., Achouri, Y., Dauguet, N., Rosa, D. R., Gunning, P. T., Manfroid, I., Lemaigre, F. P. (2018). “MiR-337-3p balances hepatobiliary gene expression and controls the transcriptional dynamics during hepatic cell differentiation” Hepatology, 67, 313-327.
Wingelhofer B., Neubauer H. A., Valent P., Han X., Constantinescu S.N., Gunning P.T., Müller M., and Moriggl R. (2018). “Implications of JAK-STAT signaling on gene regulation and chromatin remodeling in cancer.” Leukemia, 32, 1713-1726.
Geletu, M., Mohan, R., Arulanandam, R., Berger, A., Nabi I., Gunning P. T. and Raptis, L. (2018). “Reciprocal regulation of the Cadherin-11/Stat3 axis by caveolin-1 in mouse fibroblasts and lung carcinoma cells.” BBA Molecular Cell Research, 5, 1865.
Wingelhofer B., Maurer B., Heyes E. C., Cumaraswamy A. A., Berger A., de Araujo E. D., Orlova A., Freund P., Ruge F., Park F., Tin G., Ahmar S., Lardeau C. H., Sadovnik I., Bajusz D., Keserű G. M., Grebien F., Kubicek S., Valent P., Gunning P.T.,* and Moriggl R. (2018).* “Pharmacologic inhibition of STAT5 in acute myeloid leukemia.” Leukemia, 32, 1135.
Orlova A., Wingelhofer B., Neubauer H. A., Maurer B., Berger A., Keserű G. M., Gunning P. T., Valent P., and Moriggl R. (2018).* “Emerging therapeutic targets in myeloproliferative neoplasms and peripheral T cell leukemias and lymphomas.” Ex. Opin. Ther. Targets, 22, 45-57.
Murcar-Evans, B. I., Cabral, A. D., French, S., Toutah, K., De Araujo, E. D., Lai, A., Macdonald, P. M., Brown, E. D., Kraskouskaya, D., and Gunning, P. T. (2017).* “ProxyPhos sensors for the detection and quantification of negatively charged membranes.” Analyst., 142, 4511-4521.
Gleixner, K. V., Schneeweiss, M., Herrmann, H., Blatt, K., Berger, D., Eisenwort, G., Greiner, G., Byrgazov, K., Hoermann, G., Konopleva, M., Fang, J., Cumaraswamy, A. C., Gunning, P. T., Maeda, H., Moriggl, R., Deininger, M., Lion, T., Andreeff, M., Valent, P. (2017). “Combined targeting of STAT3 and STAT5: a novel approach to overcome drug resistance in Ph+ CML.” Haematologica, 102, 1519-1529.
Cabral, A. D., Murcar-Evans, B. I., Toutah, K., Bancerz, M., Rosa, D., Yuen, K., Radu, T. B., Ali, M., Penkul, A., Kraskouskaya, D., Gunning, P. T. (2017)* “Characterization and application studies of ProxyPhos, a chemosensor for the detection of proximally phosphorylated peptides and proteins in aqueous solutions.” Analyst, 142, 3922-3933.
Niit, M., Arulanandam, R., Cass, J., Geletu, M., Gunning, P. T., Elliott, B., and Raptis, L. (2017). “Regulation of HC11 Mouse Breast Epithelial Cell Differentiation by the E-cadherin/Rac axis.” Exp Cell Res., 361, 112-125.
De Araujo, E. D., Manaswiyoungkul, P., Israelian, J., Yuena, K., Farhangia, S., Berger, A., Abu Jazar, L., Gunning, P. T. (2017).* “High-throughput thermofluor-based assays for inhibitor screening of STAT SH2 domains.” J. Pharm. Biomed. Anal., 143, 159-167.
Kraskouskaya, D., Cabral, A. D., Fong, R., Bancerz, M., Rosa, D. A., Gardiner, J. E., De Araujo, E. D., Duodu, E., Armstrong, D., Fekl, U., Gunning, P. T. (2017)* “Characterization and application studies of ProxyPhos, a chemosensor for the detection of proximally phosphorylated peptides and proteins in aqueous solutions.” Analyst,142, 2451-2459.
Paiva, S-L, da Silva, S. R., de Araujo, E., Gunning, P. T. (2017).* “Regulating the Master Regulator: Controlling Ubiquitination by Thinking Outside the Active Site.” J. Med. Chem., 61, 405-421.
Garg, N., Bakhshinyan, D., Venugopal, C., Rosa, D. A., Vijayakumar, T., Manoranjan, B., Hallett, R., McFarlane, N., Mahendram, S., Delaney, K., Kwiecien, J., Arpin, C. C., Lai, P-S., Gomez-Biagi, R. F., Ali, A. M., Ajani, O. A., Hassell, J., Gunning, P. T., Singh, S. K. (2017). “CD133+ brain tumour initiating cells are dependent on STAT3 signaling to drive medulloblastoma recurrence.” Oncogene, 5, 606-617.
de Araujo, E. D., Geletu, M., Gunning, P. T. (2017).* “Strategies for over-expression and purification of recombinant full length STAT5B in Escherichia coli.” Protein Expr. Purif., 129, 1-9.
Gottardt, D., Maurer, B., Nivarthi, H., Wingelhofer, B., Prchal-Murphy, M., Chen, D., Winkler, S., Merkel, O., Schlederer, M., Prochazkova, J., Schiefer, A-S., Gurnhofer, E., Hofbauer, M., Thi Thanh Pham, H., Hochgatterer, B., Bauer, E., Hoermann, G., Hoelbl-Kovacic, A., Cumaraswamy, A. A., Lewis, A. M., Eder, J., Kitzwoegerer, M., Han, X., Valent, P., Stoiber, D., Kolbe, T., Loizou, J. I., Grebien, F., Kenner, L., Gunning, P. T., Kralovics, R., Sexl, V., Mueller, M., Rülicke, T., Moriggl, R. (2016). “STAT5 is a key regulator in NK cells and act as molecular switch from tumor surveillance to tumor promotion.” Cancer Discovery, 4, 414-429.
Gunning, P. T.,* Kraskouskaya, D., Park, J., Cabral, A. D., Gomez-Biagi, R. F. (2016). “Fluorescence-based Chemosensors for the detection of biologically relevant Phosphates in water.’ Book chapter in ‘Comprehensive Supramolecular Chemistry II’ ISBN: 978-0-12-803199-5
Linher-Melville, K., Nashed, M. G., Ungard, R., Gunning, P. T., Singh, G. (2016). “Chronic inhibition of STAT3/STAT5 in treatment-resistant human breast cancer cell subtypes: convergence on the ROS/SUMO pathway and its effects on xCT expression and system xc- activity.” PLoSOne., 11: e0161202.
Wilson A.* and Gunning P. T. (2016).* “Understanding Protein–Protein Interactions: Essential Players in(Patho)physiology.” ChemMedChem, 11, 732-733. IF = 3.016.
Ali, A.M., Gómez-Biagi, R. F., Rosa, D. A., Lai, P-S., Heaton, W. L., Park, J., Eiring, A. M., Vellore, N. A., de Araujo, E. D., Ball, D. P., Shouksmith, A. E., Patel, A. B., O’Hare, T., Deininger, M. W., Gunning, P. T. (2016).* “Disarming an electrophilic warhead: Retaining potency in TKI-resistant CML cell lines, while circumventing pharmacokinetic liabilities.” ChemMedChem, 11, 850–861.
Belton, A., Koo, M., Huso, D., Huso, T., Xian, L., Turkson, J., Gunning, P. T.*, Resar, L. (2016). “STAT3 inhibitor has potent anti-tumor activity in B-lineage acute lymphoblastic leukemia cells overexpressing the High Mobility Group A1 (HMGA1)-STAT3 pathway.” Leukemia and Lymphoma., 11: 2681-2684.
Ball, D. P., Lewis, A. M., Williams, D., Resetca, D., Wilson, D. J., Gunning, P. T. (2016). “Signal transducer and activator of transcription 3 (STAT3) inhibitor, S3I-201, acts as a potent and non-selective alkylating agent.” Oncotarget., 7, 20669-20679.
Arpin, C. C., Mac, S., Cheng, H., Jiang, Y., Page, B. D. G., Kamocka, M. M., Haftchenary, S., Su, H., Ball, D., Rosa, D., Lai, P-S., Gómez-Biagi, R. F., Ali, A. M., Kerman, K., Fishel, M. L., and Gunning, P. T. (2016). “Applying Small Molecule Signal Transducer and Activator of Trancription-3 (STAT3) Protein Inhibitors as Pancreatic Cancer Therapeutics.” Mol. Cancer Ther., 5, 794-805.
da Silva, S. R., Paiva, S-L., Bancerz, M., Geletu, M., Lewis, A. M., Chen, J., Cai, Y., Lukkarila, J. L., Dhe-Paganon, S., Li, H., Gunning, P. T. (2016). “A selective inhibitor of the UFM1-activating enzyme, UBA5.” BMCL, 26, 4542-4547.
Duodu, E., Kraskouskaya, D., Gunning, P. T. (2016). “A tool for the selective sequestration of ATP and PPi to aid in-solution phosphopeptide detection assays.” Analyst, 141, 820 – 822.
Shouksmith, A. E., Gunning, P. T. (2016). “Targeting Signal Transducer and Activator of Transcription (STAT) 3 with Small Molecules.” Book chapter in ‘Small Molecule Transcription Factors in Oncology.’ by Royal Society of Chemistry Book, ISBN: 9781788015271
Lai, P.L., Rosa, D. A., Ali, A. M., Gomez-Biagi, R. E., Ball, D. P., Shouksmith, A. E., Gunning, P. T. (2015). “A STAT inhibitor patent review: progress since 2011.” Ex. Opin. Ther. Pat., 12, 1397-421.
Haftchenary, S, Jouk, A. O., Aubry, I., Lewis, A. M., Landry, M., Ball, D. P., Shouksmith, A. E., Collins, C. V., Tremblay, M. L.,* Gunning, P. T. (2015).* “Identification of Bidentate Salicylic Acid Inhibitors of PTP1B” ACS Med. Chem. Lett., 6, 982–986.
Singh, M., Garg, N., Venugopal, C., Hallett, R., Tokar, T., McFarlane, N., Arpin, C. C., Page, B. D. G., Haftchenary, S., Todic, A., Rosa, D. A., Lai, P-S., Gómez-Biagi, R., Ali, M., Lewis, A., Geletu, M., Mahendram, S., Bakhshinyan, D., Manoranjan, B., Vora, P., Qazi, M., Murty, N. K., Hassell, J. A., Jurisica, I., Gunning, P. T., Singh, S. K. (2015). “STAT3 pathway regulates lung-derived brain metastasis initiating cell capacity through miR-21 activation.” Oncotarget 2015, 6, 27461-27477.
Linher-Melville, K., Haftchenary, S., Gunning, P. T., Singh, G.(2015).* “Signal transducer and activator of transcription 3 and 5 regulate system Xc- and redox balance in human breast cancer cells.” Mol. Cell. Biochem., 405, 205-221.
Lewis, A. M., Rana, R., Park, J-S., Gomez, R., Shaheen, A., Rosa, D., Gunning, P. T. (2015). “Developing inhibitors of STAT proteins: Targeting downstream of the kinases for treating disease.” Book chapter in “Kinomics: Approaches and Applications” by Wiley-VCH, ISBN: 978-3-527-33765-1
Zeidan, N., Su, H., Aitken, M., Gunning, P. T., Kerman, K. (2015).* “Magnetic bead-based electrochemical detection of interaction between epigallocatechin-3-gallate and STAT proteins.” Anal. Methods, 7, 3566-3569.
Arabzadeh, A., Dupaul-Chicoine, J., Breton, V., Haftchenary, S., Yumeen, S., Turbide, C., Saleh, M., McGregor, K., Greenwood, C. M. T., Akavia, U. D., Blumberg, R. S., Gunning, P. T., Beauchemin, N. (2015). “Carcinoembryonic Antigen Cell Adhesion Molecule 1 long isoform modulates malignancy of poorly differentiated colon cancer cells.” Gut, 65, 821-829.
Duodu, E., Kraskouskaya, D., Campbell, J., Graca-Lima, G., Gunning, P. T. (2015). “Selective detection of tyrosine-containing proximally phosphorylated motifs using an antenna-free Tb3+ luminescent sensor.” ChemComm, 51, 6675 - 6677.
Eiring, A. M., Page, B. D. G., Kraft, I. L., Zhang, T. Y., Vellore N. A., Reynolds, K. R., Senina, A., Pomicter, A. D., Khorashad, J. S., Gu, Z., Anderson, D. J., Zabriskie, M. S., Arpin, C. C., Cologouri, R., Ahmad, S., Moriggl, R., Baron, R., O’Hare, T., Gunning, P. T.,* Deininger, M. W. (2015).* “Combined STAT3 and BCR-ABL1 Inhibition Induces Synthetic Lethality in Therapy-Resistant Chronic Myeloid Leukemia.” Leukemia, 29, 586-597.
