
Communications Biology publishes Dr. Wassim Elkhatib's paper
Please join us in congratulating Dr. Wassim Elkhatib, PhD, Senatore lab, on his paper: Function and phylogeny support the independent evolution of an ASIC-like Deg/ENaC channel in the Placozoa published by Communication Biology.
https://www.nature.com/articles/s42003-023-05312-0#citeas
https://doi.org/10.1038/s42003-023-05312-0
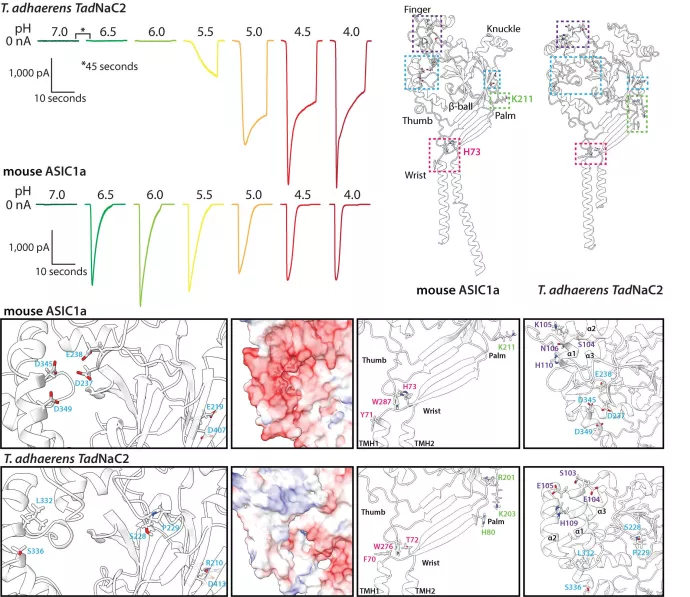
Degenerin/epithelial sodium channels (Deg/ENaC channels) were first discovered in the nematode worm C. elegans as the degenerin channels, which are gated by mechanical stimuli and involved in touch perception. Subsequently, degenerin channels were found to belong a large and phylogenetically diverse family, with an equally diverse set of activating ligands beyond mechanical stimuli including extracellular protons (i.e., pH sensing), neuropeptides (neurotransmission), and bile acids. Mammals possess three types of Deg/ENaC channels: the epithelial sodium channels which are leak sodium channels expressed in lung and kidney epithelia involved in osmoregulation, the bile-acid sensitive ion channels (BASICs) which are involved in digestion, and the acid-sensing ion channels (ASICs) which are expressed in the central and peripheral nervous system where they play important roles in extracellular H+/pH sensing, for example at synapses where fusion of synaptic vesicles results in acidification of the extracellular environment. Interestingly, more recent work has shown that ASIC channels are of common descent having diverged from other Deg/ENaC channels near the base of the Bilateria (i.e., animals with bilateral body symmetry). Notably, there are also several reports of other Deg/ENaC channels being sensitive to protons, for example in C. elegans, the fruit fly Drosophila melanogaster, and the sea anemone Nematostella vectensis, however whether these channels evolved this capacity independently from ASIC channels has not been thoroughly explored.
In our study led by PhD student Wassim Elkhatib recently published in the journal Communications Biology (6, 951 (2023). https://doi.org/10.1038/s42003-023-05312-0 ), we discovered that the early diverging non-bilaterian animal Trichoplax adhaerens, a simple sea water animal with only six cell types and lacking a nervous system, possesses a proton-activated Deg/ENaC channel with striking functional resemblance to mammalian ASIC channels. Through comparative sequence and structural analysis, we found that the Trichoplax channel, named TadNaC2, lacks all known molecular determinants for proton activation of ASIC channels. Instead, mutation analysis revealed that the Trichoplax channel harbours unique amino acids within key structural regions that are important for its proton activation. By conducting a thorough phylogenetic analysis of the Deg/ENaC family, we found that TadNaC2 is not phylogenetically-related to ASIC channels, but rather, to the mammalian BASIC channels. Together, our functional and phylogenetic analyses provide compelling evidence for the independent evolution of proton-activated Deg/ENaC channels in Trichoplax, and the most detailed functional characterization of a non-ASIC proton-activated channel to date. Furthermore, our phylogenetic analyses uncovered numerous novel clades of Deg/ENaC channels within animals, and the first phylogenetic evidence for the existence of Deg/ENaC channels outside of animals in select lineages of single-celled eukaryotes.
1st author Dr. Wassim Elkhatib Bio:
I am a recent Ph.D. graduate from the Senatore Lab at the University of Toronto Mississauga. I have always been fascinated by biological systems and the complexity of life. Indeed, even in organisms that may appear relatively simple when compared to more complex animals, evolution has sculpted intricate and sophisticated biological systems. My Ph.D. explored the evolution of Degenerin/Epithelial Sodium channels (DEG/ENaCs) and their functional properties in early diverging animals. Our research focused on an intriguing invertebrate species, Trichoplax adhaerens, which diverged from us over 600 million years ago. My journey now continues as a postdoctoral fellow in Dr. Van Petegem's lab at UBC, where I am focusing on elucidating ion channel structures using advanced Cryogenic electron microscopy and X-ray crystallography techniques. I am hoping to continue my career in academia and plan on combining electrophysiology and structural biology to understand ion channel properties, gating mechanisms, and evolution. Explore our paper for a deeper understanding of the fascinating world of ion channel evolution!
Collaborations/acknowledgements:
We would like to thank our excellent collaborators Dr. Mark A. Currie who contributed to our structural analysis of the TadNaC2 channel, Dr. Luis A. Yanez-Guerra who contributed to our bioinformatics/phylogenetic analysis of Deg/ENaC channels, and additional members of the Senatore lab who participated in the research.
