
UTM alumnus leads team working to identify potential drugs to target SARS-CoV-2
Developing drug therapies and a vaccine is high priority now as COVID-19 impacts lives around the globe.
A team of applied scientists at Cyclica Inc. led by Vijay Shahani, a U of T Mississauga alumnus, has identified thousands of small molecule drugs that can potentially be repurposed to target the SARS-CoV-2 virus that causes COVID-19.
Of the almost 10,000 drugs they evaluated through machine learning and informatics, nearly 4,500 are either approved medicines already in use or experimental medicines, with associated safety data, currently being tested in phase II and phase III clinical trials.
“The key here is speed,” says Shahani, who earned a PhD in medicinal chemistry from UTM before returning to earn a Master of Science in Biomedical Communications.
Andrew Haller, an adjunct lecturer in the Department of Pharmacology and Toxicology at the University of Toronto, says a resource of approved drugs could potentially expedite COVID-19 treatments by months or even years.
"Using an already approved therapeutic would reduce the regulatory requirements to get into the clinic. On-market drugs have a lot of data associated with them and people understand the risks and methods of use of the drugs. You could deliver new therapeutics to battle COVID-19 in as little as a few months," says Haller, who is also founder and CEO of Phoenox Pharma, one of Cyclica’s local biotech partners.
Cyclica’s team has stored the collection of drugs they identified, along with protein data, in a searchable database named PolypharmDB. They developed the database using Cyclica’s deep learning engine MatchMaker, which predicts pairings of drugs to proteins.
“We used the literature surrounding infections caused by the previous viruses SARS and MERS to propose potential host proteins that may serve as drug targets for SARS-CoV-2. We then queried PolypharmDB to select small molecules that are predicted to bind to those key proteins,” says Shahani.
“The results from PolypharmDB are small molecule drug candidates that MatchMaker predicted to be likely binders to key host proteins,” he says.
MatchMaker predicted that the drug disopyramide, used to treat irregular heartbeats, will interact with a human protein involved in SARS-CoV-2 infection. “This prediction, if validated pre-clinically, could translate into an acute treatment for COVID-19."
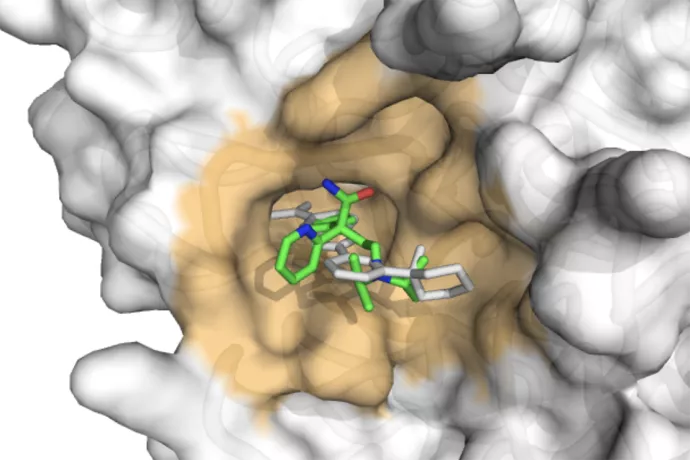
Cyclica's AI technology predicted that the antiarrythmic drug disopyramide will interact with the human protein TMPRSS2 involved in SARS-CoV-2 infection. Vijay Shahani says that this prediction, if validated pre-clinically, could translate into an acute treatment for COVID-19. (Visualization submitted by V. Shahani)
Cyclica is also offering their support to the research and academic communities working within the realm of COVID-19.
"Let's say that there is another host protein identified by researchers as critically involved with a COVID-19 infection. What we can do is very quickly mine PolypharmDB and pull out 10, 50,100 candidate drug molecules, all with safety data, that are likely to interact with that host protein. Depending on the target, that could be done in less than 24 hours," Shahani says.
Cyclica, a biotechnology company in Toronto, uses artificial intelligence to design new medicines and accelerate drug discovery efforts. Shahani says that they have had the ability to assess small molecule drugs for their off-target interactions for a while now.
"But we've applied it to smaller chemical libraries that our collaborators at U of T and Ryerson were interested in. Now, we have the chance to assess these existing medicines to see how they could make an impact on the current COVID-19 pandemic and that's what prompted us to build it out so quickly,” he says.
Researchers who have a target and a hypothesis and who wish to interact with Cyclica's technologies can complete and submit a form through their COVID-19 Response page.
"To accelerate drug discovery is our mandate. The ultimate goal would be to try and validate some of these small molecule drugs in preliminary assays and then push them forward into the clinics if we can,” says Shahani. “I think we felt that we have an opportunity here to help save people's lives."
