Aidan Levinsky, Gregor McEdwards & Nasha Sethna from Currie Lab Share Cofirst Authorship
Aidan Levinsky, Gregor McEdwards & Nasha Sethna from Currie Lab share cofirst authorship for: Targets of histone H3 lysine 9 methyltransferases published by Frontiers in Cell and Developmental Biology
https://doi.org/10.3389/fcell.2022.1026406
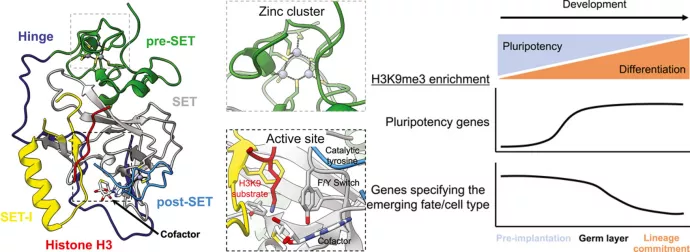
Histone H3 lysine 9 di- and trimethylation are well-established marks of constitutively silenced heterochromatin domains found at repetitive DNA elements including pericentromeres, telomeres, and transposons. Loss of heterochromatin at these sites causes genomic instability in the form of aberrant DNA repair, chromosome segregation defects, replication stress, and transposition. H3K9 di- and trimethylation also regulate cell type-specific gene expression during development and form a barrier to cellular reprogramming. However, the role of H3K9 methyltransferases extends beyond histone methylation. There is a growing list of non-histone targets of H3K9 methyltransferases including transcription factors, steroid hormone receptors, histone modifying enzymes, and other chromatin regulatory proteins. Additionally, two classes of H3K9 methyltransferases modulate their own function through automethylation. Here we summarize the structure and function of mammalian H3K9 methyltransferases, their roles in genome regulation and constitutive heterochromatin as well as the current repertoire of non-histone methylation targets including cases of automethylation.