
Sarah Rauscher
-
E-mail:
-
Phone:
-
Fax:
-
Mailing Address:
3359 Mississauga Road
Mississauga ON L5L 1C6
Canada
Research Areas:
Computational Biophysics
Research Profile:
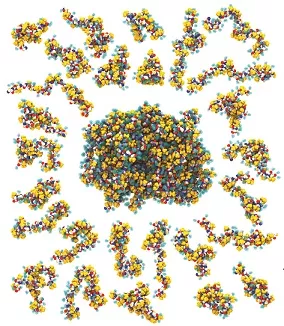
Intrinsically disordered proteins (IDPs) are abundant in all kingdoms of life and fulfill many critical biological functions. IDPs have been found, for instance, to regulate the cell cycle, transcription and translation, act as hubs in signalling networks, and maintain tight control over the locations of other proteins in the cell. IDPs also play roles, which are yet to be fully understood, in various diseases, such as Alzheimer’s, Parkinson’s, ALS, cancer, and AIDS. The tremendous biological importance of IDPs provides strong motivation for their detailed biophysical characterization.
IDPs are still poorly understood relative to the wealth of structural information available for folded proteins. IDPs are notoriously difficult to study using experimental approaches. Moreover, on a fundamental level, their description poses formidable challenges: IDPs do not have a unique structure, but rather a structural ensemble consisting of many interconverting conformational states. Classical molecular dynamics (MD) simulations, with their empirical force fields, can be used to model the ensembles of IDPs, but there remain significant challenges in both statistical sampling and accuracy. Our previous research involved carrying out a systematic comparison of state-of-the-art force fields for the specific case of IDPs and developing an improved force field suitable for both IDPs and folded proteins (Rauscher et al. J. Chem. Theor. Comput. 2015, Huang et al. Nature Methods 2017).
The fact that accurate structural descriptions are still not available for many of the disordered proteins underlying human diseases severely impedes rational drug development. On a more fundamental level, we need a new structure-function paradigm that describes how the sequences of IDPs dictate their structural ensembles, and how the structural properties of IDPs give rise to their diverse cellular functions. Research in our group addresses this gap in our knowledge by obtaining atomistic descriptions of the structure and dynamics of IDPs. Comparison to experimental data is an integral part of our research program, not only to validate the structural ensembles, but also to improve parameters specifically for IDPs. The long-term goal of these studies is to develop a simulation methodology to obtain structural descriptions of IDPs as efficiently and accurately as possible. Methodological insights gained from studying specific IDPs will drive the continuous improvement of a robust simulation methodology for the study of IDPs in general.
There are currently open positions in the group for graduate and undergraduate students. Interested candidates should send inquiries by email to sarah.rauscher@utoronto.ca.
Courses Taught:
Publications
Klyshko, E., Kim, J. S., Rauscher, S. (2023) LAWS: Local alignment for water sites — Tracking ordered water in simulations. Biophysical Journal (in press). doi:10.1016/j.bpj.2022.09.012
Armstrong, H. K., Bording-Jorgensen, M., Santer, D. M., Zhang, Z., Valcheva, R., Rieger, A. M., Kim, J. S., Dijk, S. I., Mahmood, R., Ogungbola, O., Jovel, J., Moreau, F., Gorman, H., Dickner, R., Jerasi, J.,Mander,I. K., Lafleur, D., Cheng, C., Petrova, A., Jeanson, T. L., Mason, A., Sergi, C. M., Levine, A., Chadee, K., Armstrong, D., Rauscher, S., Bernstein, C. N., Carroll, M. W., Huynh, H. Q., Walter, J., Madsen, K. L., Dieleman, L. A., Wine, E. (2023) Unfermented β-fructan fibers fuel inflammation in select inflammatory bowel disease patients. Gastroenterology 164, 228–240 doi:10.1053/j.gastro.2022.09.034
Jang, J., Reed, P. M. M., Rauscher, S., Woolley, G. A. (2022) Point (S-to-G) Mutations in the W (S/G) GE Motif in Red/Green Cyanobacteriochrome GAF Domains Enhance Thermal Reversion Rates. Biochemistry 61, 1444-1455. doi:10.1021/acs.biochem.2c00060
Lazar, T., Guharoy, M., Vranken, W., Rauscher, S., Wodak, S. J., & Tompa, P. (2020). Distance-Based metrics for comparing conformational ensembles of intrinsically disordered proteins. Biophysical Journal, 118(12), 2952-2965.
McGough, L., Kim, J., Klyshko, E., Ranganathan, R., & Rauscher, S. (2020). Evaluating molecular simulations of protein dynamics using novel experimental data. Bulletin of the American Physical Society, 65.
Kim, J. S., McGough, L., Klyshko, E., Ranganathan, R., & Rauscher, S. (2020). Novel Simulation Methods for Electric Field-Induced Dynamics of Protein Crystals. Biophysical Journal, 118(3), 142a.
Klyshko, E., McGough, L., Kim, J. S., Ranganathan, R., & Rauscher, S. (2020). EF-X in Silico-Modeling Protein Dynamics in an Electric Field. Biophysical Journal, 118(3), 504a.
Meneksedag-Erol, D., de Araujo, E. D., Erdogan, F., Seo, H. S., Dhe-Paganon, S., Gunning, P. T., & Rauscher, S. (2020). Cancer Activating Mutations in STAT5B: Elucidating the Impact on Protein Structure and Dynamics using Atomistic Molecular Simulations. Biophysical Journal, 118(3), 504a.
Haas-Neill, L. I., & Rauscher, S. (2020). Molecular Dynamics Simulations of the A2A G-Protein Coupled Receptor. Biophysical Journal, 118(3), 524a.
Kim, J. S., & Rauscher, S. (2019). Hamiltonian Replica Exchange for Enhanced Sampling of the Conformational Landscape for Intrinsically Disordered Proteins. Biophysical Journal, 116(3), 143a-144a.
Klyshko, E., & Rauscher, S. (2019). Defining Conformational States of Proteins using Dimensionality Reduction and Clustering Algorithms. Biophysical Journal, 116(3), 290a.
Meneksedag-Erol, D., & Rauscher, S. (2019). Atomistic Simulation Tools to Study Protein Self-Aggregation. In Protein Self-Assembly (pp. 243-262). Humana, New York, NY.
Bandyopadhyay, A., Van Eps, N., Eger, B. T., Rauscher, S., Yedidi, R. S., Moroni, T., ... & Ernst, O. P. (2018). A novel polar core and weakly fixed C-tail in squid arrestin provide new insight into interaction with rhodopsin. Journal of molecular biology, 430(21), 4102-4118.
