
David R. McMillen
-
E-mail:
-
Phone:
-
Fax:
-
Mailing Address:
3359 Mississauga Road
Mississauga ON L5L 1C6
Canada
Research Areas:
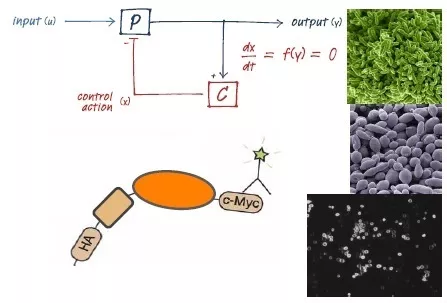
Synthetic biology, systems biology, cellular dynamics
The Synthetic Biology and Cellular Control Laboratory seeks to create an engineering discipline within living cells. We use molecular biology to design and experimentally implement biological “devices” that sense the internal states of cells, and respond to those states with outputs that change the cell’s dynamics in useful ways (implementing feedback control loops to regulate a protein’s expression level, for example). We are using our interdisciplinary expertise to translate abstract mathematical results from fields like control theory into practical design procedures to be used in the context of biology. We also have projects on expanding the library of available parts to be used in biological designs, including harnessing novel forms of biological regulation (like activating RNA) and developing new forms of regulatory targeting (like an engineered polymerase that can initiate transcription wherever desired).
The group has a strong and growing interest in the applied results of our research, particularly in the area of global health. Current projects include inexpensive cell-based biosensors and protein production systems to be used in the developing world.
Courses Taught:
JCP221H5, JCP321H5, and JCP410H5 (undergraduate); CHM1448H1 (graduate)
Publications
Criselda Jean G. Cruz, Richard Kil, Stanley Wong, Louis C. Dacquay, Denise Mirano-Bascos, Pilarita T. Rivera, and David R. McMillen (2022). Malarial antibody detection with an engineered yeast agglutination assay. ACS Synthetic Biology 11: 2938-2946. https://doi.org/10.1021/acssynbio.2c00160
Louis C. Dacquay and David R. McMillen (2021). Improving the design of an oxidative stress sensing biosensor in yeast. Federation of European Microbiological Societies (FEMS) Yeast Research. (https://doi.org/10.1093/femsyr/foab025).
Milos Legner, David R. McMillen, and Dennis G. Cvitkovitch (2019). Role of dilution rate and nutrient availability in the formation of microbial biofilms. Frontiers in Microbiology. (https://doi.org/10.3389/fmicb.2019.00916).
Brendan J. Hussey and David R. McMillen (2018). Programmable T7-based synthetic transcription factors. Nucleic Acids Research 46(18): 9842–9854.
Edouard A. Harris, Alla Buzina, Jason Moffat, and David R. McMillen (2017). Design and experimental validation of small activating RNAs targeting an exogenous promoter in human cells. ACS Synthetic Biology 6(4): 628-637.
Zhe F. Tang and David R. McMillen (2016). Design principles for the analysis and construction of robustly homeostatic biological networks. Journal of Theoretical Biology 408: 274-289.
Edouard A. Harris, Eun Jee Koh, Jason Moffat, and David R. McMillen (2016). Automated inference procedure for the determination of cell growth parameters. Physical Review E 93: 012402.
Mostafizur Mazumder, Katherine E. Brechun, Yongjoo B. Kim, Stefan A. Hoffmann, Yih Yang Chen, Carrie-Lynn Keiski, Katja M. Arndt, David R. McMillen and G. Andrew Woolley (McMillen and Woolley, co-senior authors) (2015). An E. coli system for evolving improved light-controlled DNA-binding proteins. Accepted (June 2015) in Protein Engineering, Design, and Selection.
